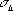 ) to get p(F; FC, D,
) to get p(F; FC, D, 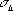 )
)Target Function vs. Optimization Method
Distinction is sometimes lost, but the two can generally be chosen independently of each other
Target Function = mathematical score for judging the fit of the model to the data
LSQ (RESI): old-fashioned least-squares target
MLF1: maximum likelihood target based on amplitudes, assuming a Gaussian error in measurement of F
MLF2: maximum likelihood target based on intensities, assuming a Gaussian error in measurement of F2
MLHL: maximum likelihood target based on amplitudes plus prior phase information encoded by Hendrickson-Lattman coefficients
Optimization method = mathematical algorithm for improving the target function by changing the model
Monte Carlo: random search using only function values
Genetic algorithms: random search using only function values
Steepest descents: search for minimum follows vector of first derivatives of target function with respect to model parameters
Simulated annealing: first derivatives of target function contribute to forces in molecular dynamics run
Conjugate gradients: second derivative information is inferred from the history of the refinement run
Gradient/curvature: uses first and unmixed second derivatives
Full-matrix: uses first derivatives and full-matrix of mixed second derivatives
Probability distribution of amplitude FO
Integrate out unknown phase from p(F; FC,
D, 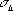 ) to get p(F; FC, D,
) to get p(F; FC, D, 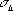 )
)
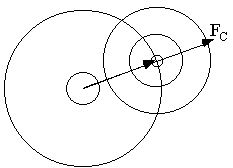
Add observational error to amplitude (or intensity)
p(FO; FC, D, 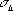 ,
,
 F) = p(F; FC, D,
F) = p(F; FC, D,
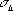 )
) 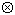 p(FO-F)
p(FO-F)
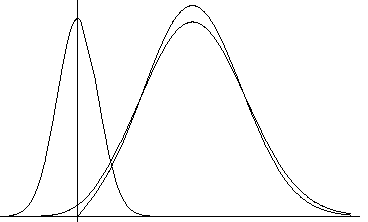
There is a good Gaussian approximation for the case of a Gaussian error in amplitudes (basis of MLF1 target):
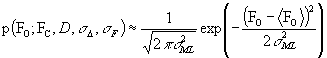
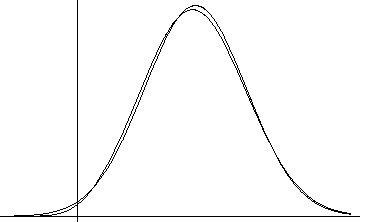
 ML incorporates contributions
of both
ML incorporates contributions
of both 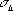 and
and  F
F
Calibrating the Likelihood Functions
The likelihood functions include the parameters 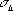 and D, each of which is a resolution-dependent parameter that can be
calculated from
and D, each of which is a resolution-dependent parameter that can be
calculated from ![]()
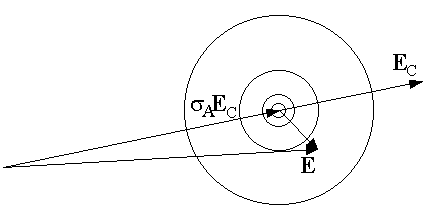
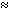 fraction of normalized
structure factor that is correct
fraction of normalized
structure factor that is correct
combined effect of errors and model incompleteness
adjust ![]() to maximize
likelihood function in SIGMAA
to maximize
likelihood function in SIGMAA
The relative weight between the X-ray terms (data likelihood) and the geometry terms (restraints) is also important
in theory, this falls out directly from respective probabilities
in practice, the diffraction data are overfit, and convergence requires overweighting the data likelihood component
the amount of overfitting depends on the resolution
so does the necessary overweighting
heuristic relative weighting is achieved either by comparing the X-ray and geometry gradients, or by trial and error
divide X-PLOR suggested weight by about two
start from about 50 for TNT
Problems in estimating 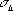
Could run original SIGMAA, then estimate 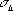 using
using 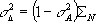
but ![]() is overestimated
with data used in refinement
is overestimated
with data used in refinement
Could use cross-validation (Rfree) data to estimate
![]()
but original SIGMAA requires 500-1000 reflections per resolution shell to get reliable estimates
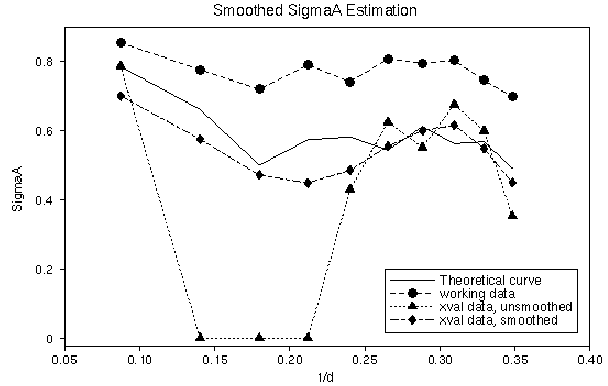
Cure: add a smoothing term to exploit the fact that
![]() values should vary smoothly with resolution.
values should vary smoothly with resolution.
this is sufficient to make the ![]() estimation behave with about 500-1000 reflections in total.
estimation behave with about 500-1000 reflections in total.
Initially Estimating and Updating ![]() Values
Values
The likelihood target depends on the validity of the ![]() values
values
must not be contaminated by refinement bias
if refinement has been carried out before setting aside cross-validation data, the overfitting bias must be removed
e.g. carry out a run of room-temperature dynamics
![]() values are estimated
in resolution bins
values are estimated
in resolution bins
there should be enough reflections in each bin to obtain numerical stability
there should be enough bins to capture the variation of
![]() with resolution
with resolution
good compromise -> set the number of bins to:
As refinement progresses, the model improves (we hope),
so ![]() values should increase
values should increase
The change in ![]() values changes the refinement target in a basic way, changing even the optimal
value of Fc
values changes the refinement target in a basic way, changing even the optimal
value of Fc
unlike the case of the least-squares target
As a result, ![]() values
should be updated once or twice through a refinement run
values
should be updated once or twice through a refinement run
Effect of Non-Crystallographic Symmetry
In the presence of NCS, randomly chosen cross-validation reflections are not independent of the working data
statistical relationships between reflections related by the rotational part of the NCS operation
depends on number of NCS-related molecules and on relationship between NCS and crystallographic symmetry
Overfitting of the working data will spread into the cross-validation set
not generally very serious for, say, 2-fold NCS
20-fold NCS hindered ML refinement of one structure we determined
Possible solution: select cross-validation data in thin resolution shells
all reflections related by rotational operations will be in the same resolution shell, so overfitting should not propagate